Rebel with a Cause
By Whitney Heins.

Thanks to Scott Emrich’s rebellious nature, the world is getting a big step closer to saving thousands of lives a year.
When the associate professor of electrical engineering and computer science—newly arrived by way of the University of Notre Dame—started his undergraduate degree, it was assumed by friends and family he would go to medical school. The thing was, he really loved working with computers—his so-called hobby in high school.
So, he went rogue and pursued bioinformatics and computational biology instead.
The result? A researcher who now has four active awards from the National Institutes of Health (NIH). His most recently funded effort is focused on conducting groundbreaking work in understanding genes in the malaria parasite that are associated with drug resistance—with the ultimate goal of reducing and eliminating the often-deadly disease.
According to World Health Organization, malaria affects roughly 215 million people a year and kills upwards of a half million. How, in 2018, are so many people are still suffering from this ancient disease? It’s because malaria increasingly is mutating to outwit modern drugs.
“The challenge is that the parasite becomes resistant and it is doing it faster and faster. First it took hundreds of years, then tens, then even faster,” explained Emrich.
Emrich is collaborating with researchers in South Bend, Seattle, and San Antonio on an NIH multi-million-dollar project that aims to uncover resistance mechanisms by conducting experimental genetic crosses of the malaria parasite. And, then analyzing massive amounts of digital data from each of these crosses.
A genetic cross is the result of breeding two different individuals, for instance, one parasite known for drug sensitivity and one known for drug resistance. The resulting offspring then carry part of the genetic material from each parent parasite, allowing researchers to identify the genes’ underlying drug resistance with enough corresponding data.
In the past, researchers used chimpanzees. But this method was costly, timely, and ethically compromising, causing NIH to halt all chimp-driven research. This time, they’re harnessing the power of mice which brings in a lot more data with a lot less time and money.
“The secret sauce of this research are these mice that can tolerate being injected with human liver and red blood cells,” explained Emrich. “The human cells grow on the mouse’s liver and this allows for rapid and routine generation of large numbers of parasite progeny.”
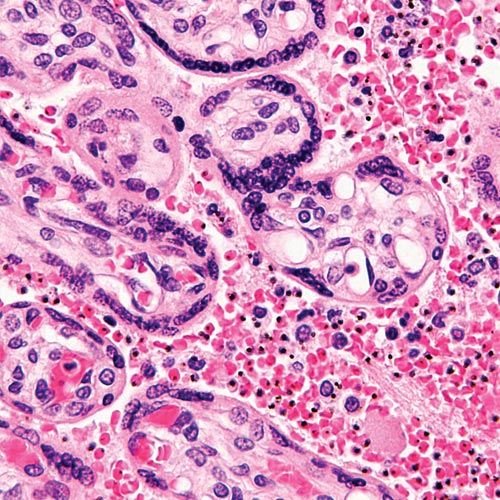
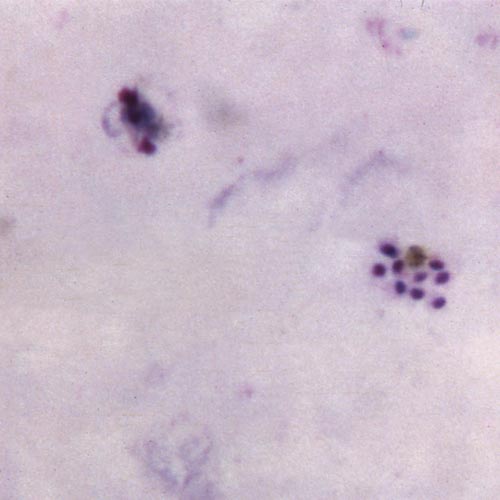
Using computational methods, Emrich can then mix and match haploids—single sets of unpaired chromosomes—in the DNA sequence to find out which genes underlie resistance.
The mouse method has already generated more genetic crosses in the past three years than in the decades prior.
“This method puts us in the position to catch drug resistance as it is happening, as well as test unknown effects of resistant Asian and deadly African hybrids in the lab, so we can come up with a plan to combat the parasite head on.”
—Scott Emrich
But malaria prevention isn’t the only area of focus for Emrich. In fact, his passions are seemingly unending.
“At Notre Dame, my colleagues joked that I worked with every biologist on campus,” shared Emrich. “But what I see as unique about bioinformatics is that we are able to help solve many problems and have the ability to train students with diverse talents through many cool projects.”
Emrich noted he has graduated nine PhD students and they were the ones who did most of the work on all his very diverse projects.
Only here since the beginning of the spring semester, Emrich is already in talks about collaborating with colleagues across UT’s campus in ecology, microbiology, and agriculture, in addition to those within his own department. Most of this early work will be collaborative with four EECS undergrads who have already joined his group meetings.
As he builds his program at UT, Emrich hopes to illuminate for his students the potential for collaboration that lies within computing.
“It can apply to so many disciplines and there are so many different career paths. I’ve had students go on to work at companies like Google, Amazon, and more traditional bio/pharma companies,” he said.
In effect, Emrich hopes his proclivity to push boundaries and rebel against the expected will open a world of possibilities—and fewer problems—for his students.